

All-new personal structured illumination super-resolution system for the individual lab.
Introducing an all-new super-resolution system for the individual lab. Achieve 2x resolution at half the price!
Utilizing structured illumination microscopy (SIM) technology, the all-new N-SIM E realizes double the spatial resolution of conventional optical microscopes (to approximately 115 nm). N-SIM E is a streamlined, affordable super-resolution system supporting only essential, commonly used excitation wavelengths and imaging modes, making it an obvious choice for individual labs.
The N-SIM E utilizes Nikon’s innovative approach to “structured illumination microscopy”. By pairing this powerful technology with Nikon’s renowned objectives (with unparalleled NA of 1.49) the N-SIM E nearly doubles the spatial resolution of conventional light microscopes to approximately 115 nm*, and enables detailed visualization of minute intracellular structures and their interactions.
* This value is measured FWHM of 100 nm beads exited with 488 nm laser in 3D-SIM mode.
|
Conventional widefield image |
Microtubules in B16 melanoma cell labeled with YFP
Objective: CFI Apochromat TIRF 100XC Oil (NA 1.49)
Image capturing speed: approximately 1.8 sec/frame (movie)
Reconstruction method: Slice
Photographed with the cooperation of: Dr. Yasushi Okada, Laboratory for Cell Polarity Regulation, Quantitative Biology Center, RIKEN
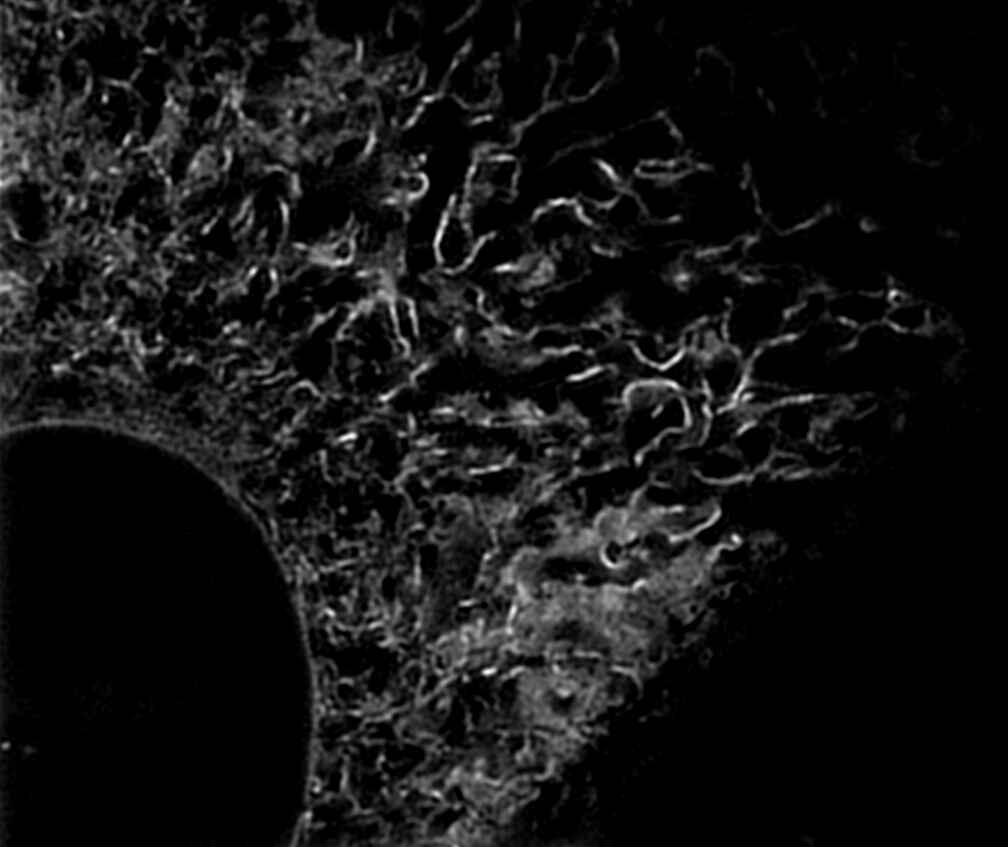
|
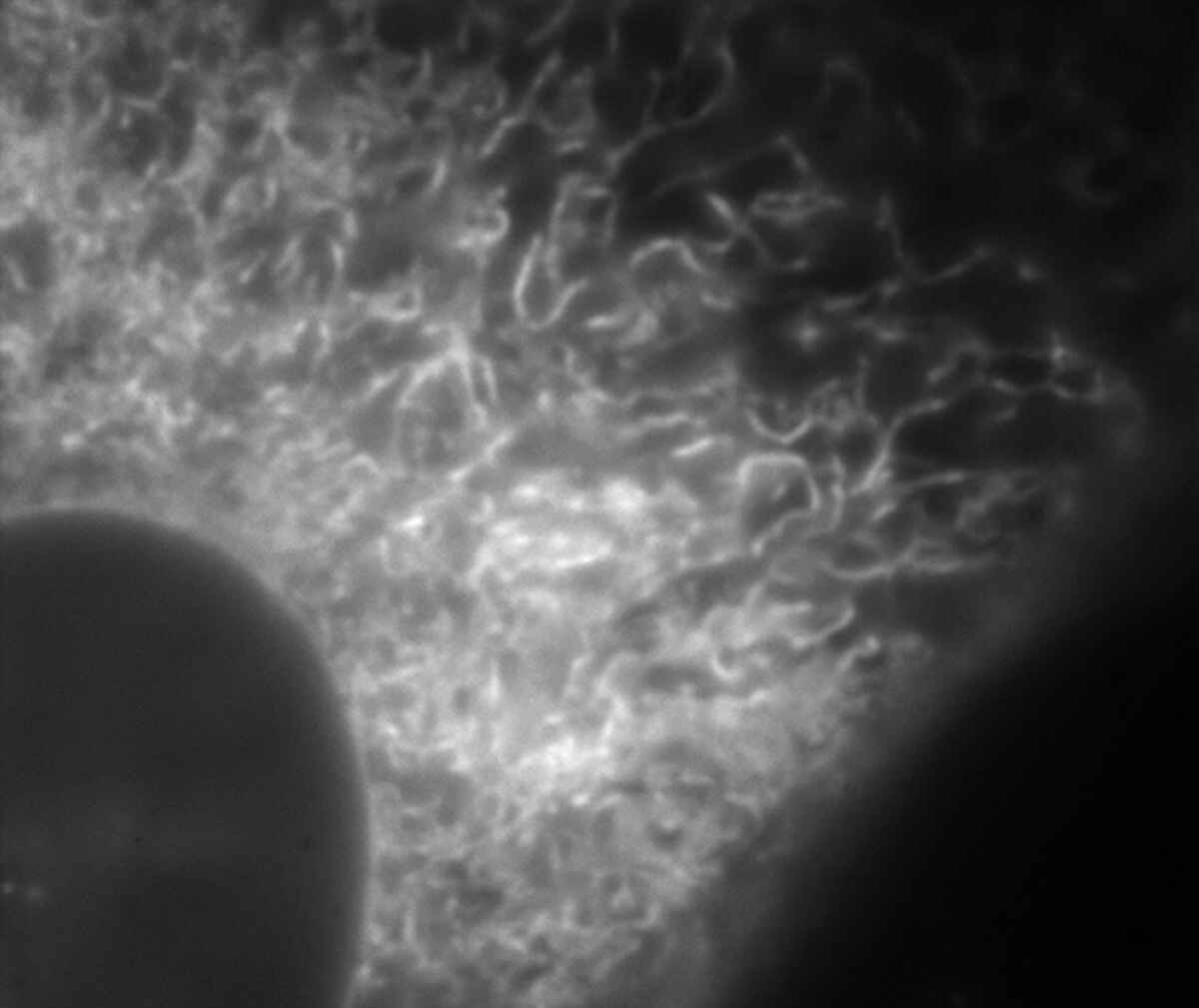
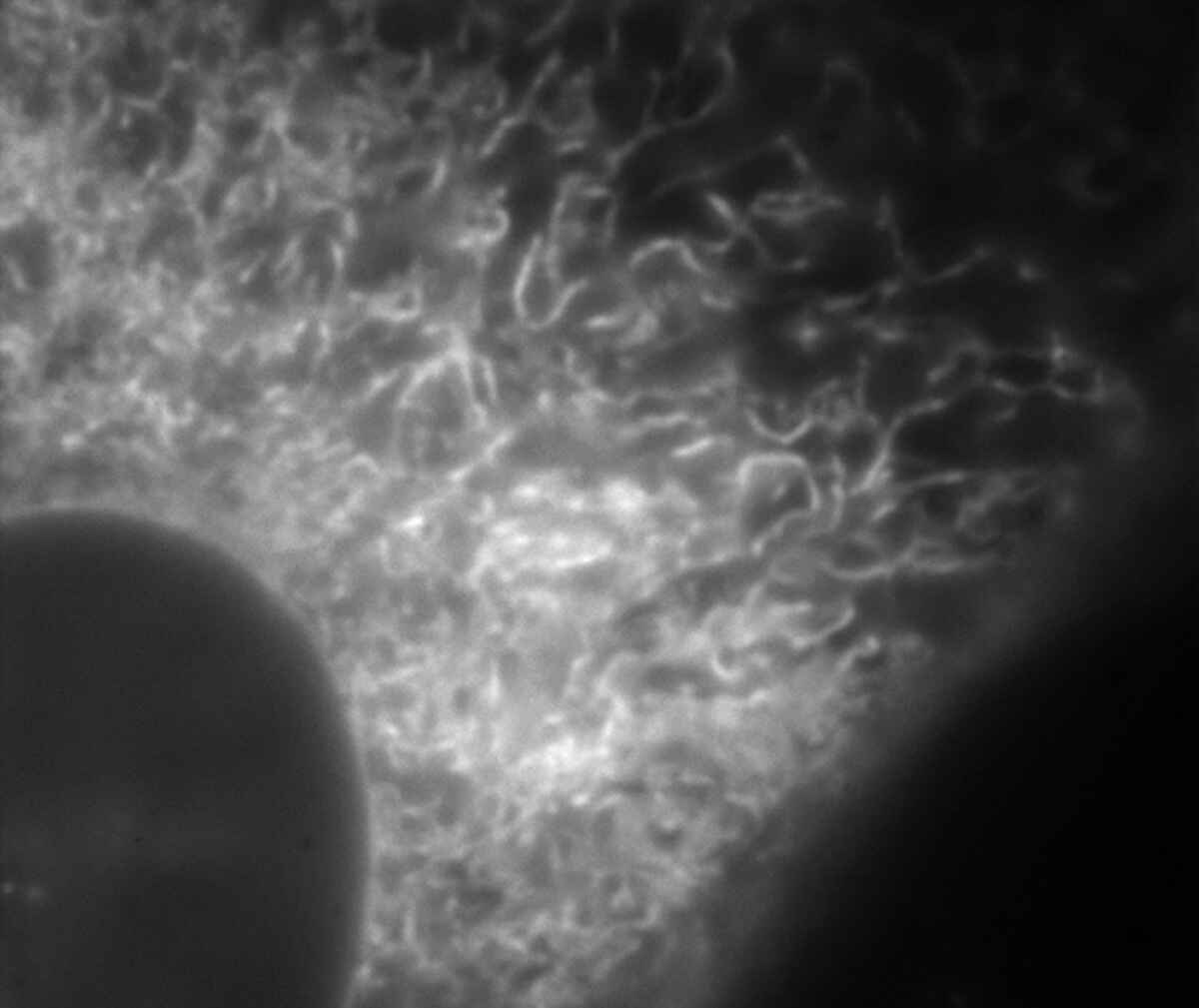
Conventional widefield image |
Endoplasmic reticulum (ER) in living HeLa cell labeled with GFP
Objective: CFI Apochromat TIRF 100XC Oil (NA 1.49)
Image capturing speed: approximately 1.5 sec/frame (movie)
Reconstruction method: Slice
Photographed with the cooperation of: Dr. Ikuo Wada, Institute of Biomedical Sciences, Fukushima Medical University School of Medicine
N-SIM E provides fast imaging performance for Structured Illumination techniques, with a time resolution of approximately 1 sec/frame, effective for live-cell imaging.
N-SIM E can acquire super-resolution images with a large field of view of 66 µm square. This larger imaging area enables very high throughput for applications/samples that benefit from larger fields of view, such as neurons, reducing the amount of time and effort required to obtain data.
Growth cone of NG108 cell labelled with TRITC-phalloidin (F-actin, orange) and Alexa Fluor® 488 (microtubules, green)
Sample courtesy of: Dr. Shizuha Ishiyama and Dr. Kaoru Katoh, The National Institute of Advanced Industrial Science and Technology (AIST)
The 3D-SIM mode generates structured illumination patterns in three dimensions to deliver a two-fold improvement in lateral and axial resolutions. Two reconstruction methods (“slice” and “stack”) are available to optimize results according to application requirements (e.g. sample thickness, speed, etc.). Slice reconstruction is suitable for imaging living cells at specific depths, as it allows axial super-resolution imaging with optical sectioning at 300 nm resolution. Optional stack reconstruction, based on Gustafsson’s theory, is suitable for acquisition of volume data as it can image thicker specimens with higher contrast than slice reconstruction.
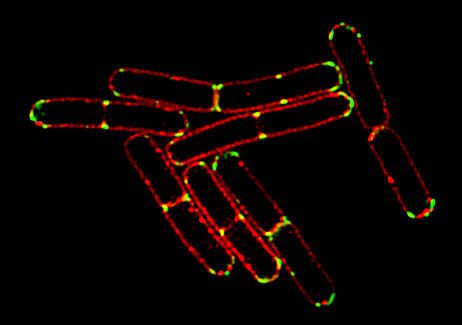 3D-SIM image |
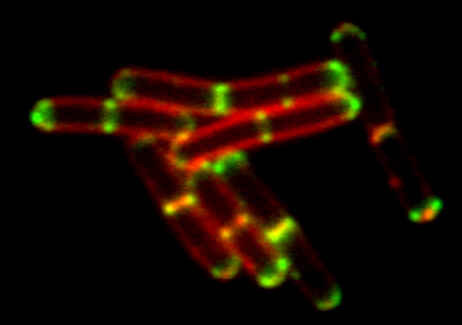
Conventional widefield image |
Bacillus subtilis bacterium stained with membrane dye Nile Red (red), and expressing the cell division protein DivIVA fused to GFP (green).
The super-resolution microscope allows for accurate localization of the protein during division.
Reconstruction method: Slice
Photos courtesy of: Drs. Henrik Strahl and Leendert Hamoen, Centre for Bacterial Cell Biology, Newcastle University
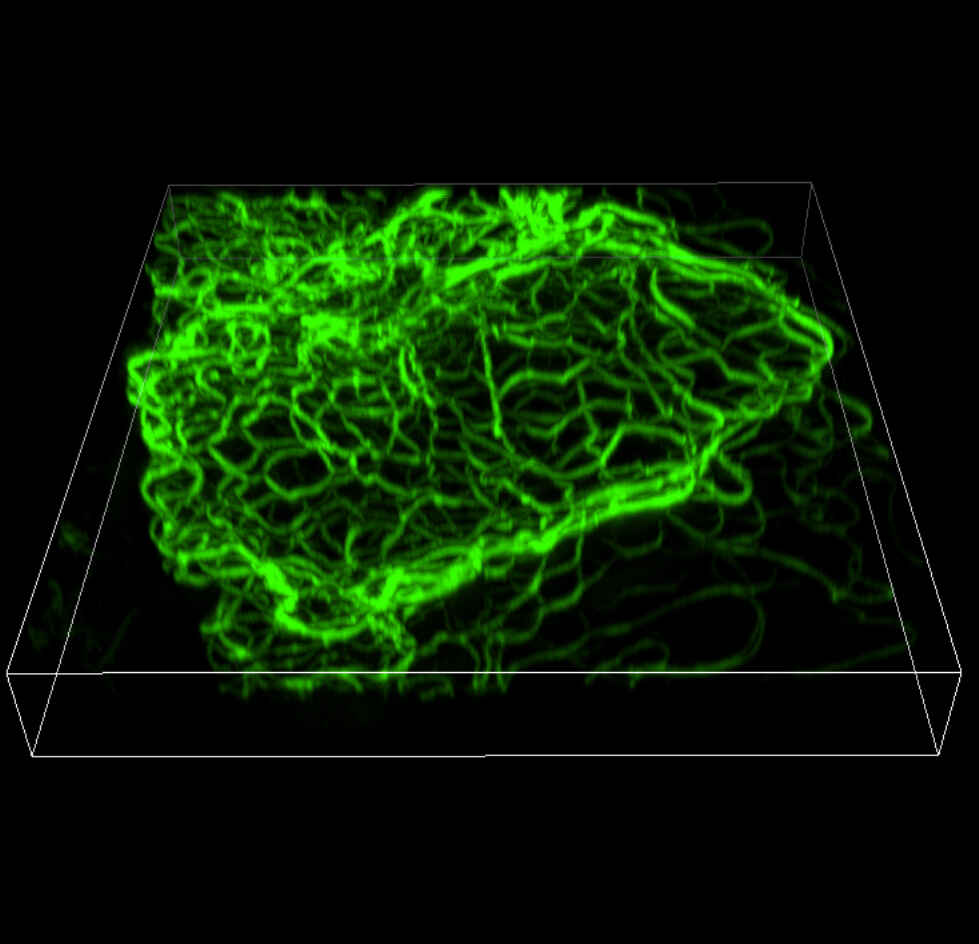 3D-SIM (Volume view)Width: 26.16 μm, Height: 27.11 μm, Depth: 3.36 μm |
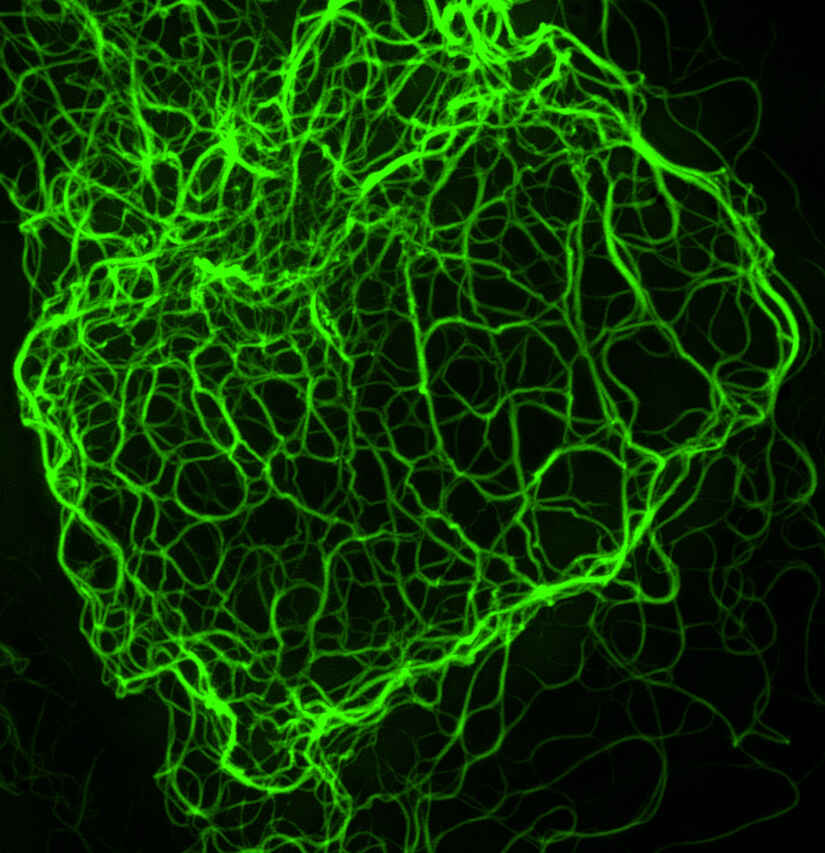 |
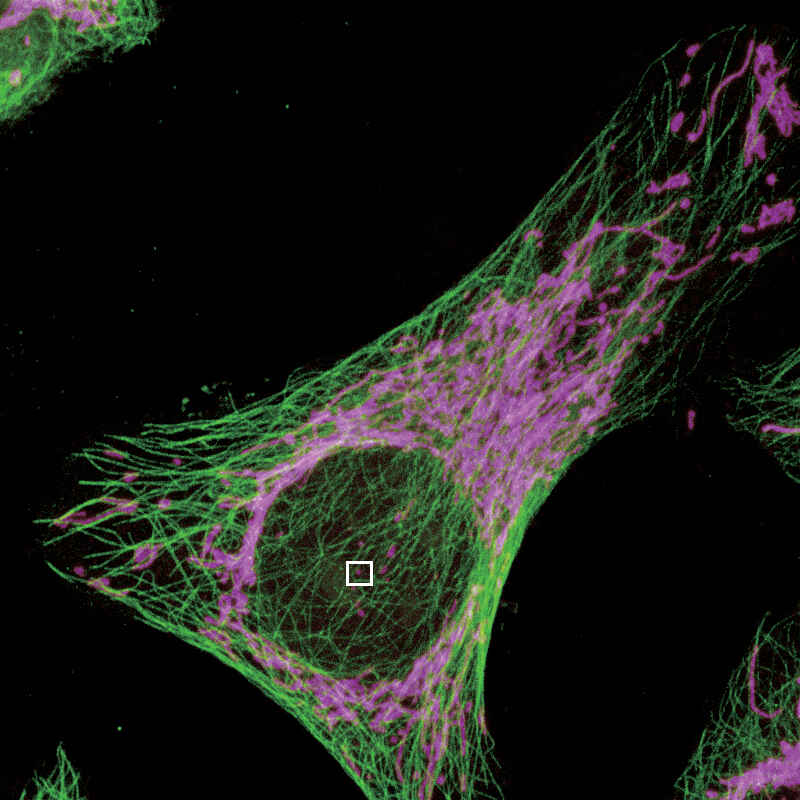
Mouse keratinocyte indirectly immunolabeled for keratin intermediate filaments and visualized with Alexa Fluor® 488 conjugated secondary antibodies.
Reconstruction method: Stack
Photo courtesy of: Dr. Reinhard Windoffer, RWTH Aachen University
The N-SIM E can be combined with a confocal microscope such as the AX/AX R. A desired location in a sample can be specified in a low-magnification/large FOV confocal image and acquired in super-resolution by simply switching the imaging method. Combining a confocal microscope with a super-resolution system can provide a method for gaining larger contextual views of super-resolution information.
 Select the location to acquire a SIM image in a confocal image |
|
The compact LU-N3-SIM laser unit dedicated for N-SIM E is installed with the three most commonly used wavelength lasers (488/561/640), enabling super-high resolution imaging in multiple colors. It enables the study of dynamic interactions of multiple proteins of interest at the molecular level.
Silicone immersion objectives use high viscosity silicone oil with a refractive index close to that of live cells as an immersion liquid. Because of this improved refractive index compatibility, these objectives can provide improved photon collection capability and resolution when performing super-resolution imaging deeper into the specimen. They exhibit superior chromatic aberration correction and high transmittance over a broad range of wavelengths.
The system can be configured with either a 100X oil immersion type, which is suitable for the imaging of fixed samples, or a 60X water immersion type, which is optimal for time-lapse live-cell imaging. The SR objectives are aligned and inspected using wavefront aberration measurement technologies to ensure the lowest possible asymmetric aberration and the superb optical performance required for super-resolution imaging.
The N-SIM E is compatible with dry objectives, making both super-resolution imaging and confocal imaging available without switching lenses. Low-magnification, wide field-of-view dry lenses enable high resolution observation even at the periphery of sample tissues.
* Dry objectives support slice reconstruction
| N-SIM E | |
|---|---|
| Lateral resolution (FWHM of beads in xy) | 115 nm*1 in 3D-SIM mode |
| Axial resolution (FWHM of beads in z) | 269 nm*1 in 3D-SIM mode |
| Image acquisition time | Up to 1 sec/frame (3D-SIM) |
| Imaging mode | 3D-SIM (Reconstruction method: slice, stack) |
| Multi-color imaging | Up to 3 colors |
| Compatible laser | LU-NV series laser unit 488 nm, 561 nm, 640 nm |
| Compatible microscope | Motorized inverted microscope ECLIPSE Ti2-EPerfect Focus System Motorized XY stage with encoders Motorized barrier filter wheel Piezo Z stage |
| Objective | CFI SR HP Apochromat TIRF 100XC Oil (NA 1.49) CFI SR HP Apochromat TIRF 100XAC Oil (NA 1.49) CFI SR Plan Apochromat IR 60XC WI (NA 1.27) CFI SR Plan Apochromat IR 60XAC WI (NA 1.27) CFI Plan Apochromat Lambda 60XC (NA 0.95)*2 CFI Plan Apochromat Lambda D 40XC (NA 0.95)*2 |
| Camera | ORCA-Fusion BT sCMOS camera (Hamamatsu Photonics K.K.) |
| Software | NIS-Elements AR NIS-Elements C (for Confocal Microscope) Both require additional software modules NIS-A 6D and N-SIM Analysis |
| Operating conditions | 20 ℃ to 28 ℃ (± 1.5 ℃) |
*1 These values are measured using 100 nm diameter beads excited by a 488 nm laser. Actual resolution is dependent on laser wavelength and optical configuration.
*2 Supports slice reconstruction.